Small, beautiful, and deadly – an influenza A virus in extreme close-up. (Centers for Disease Control and Prevention, Dan Higgins, illustrator)
By Jonas Waldenström
We live in a world of viruses. Actually, one could equally well define being alive as being infected; only when we are truly dead, we are no longer homes for any viral passengers.
Viruses are everywhere, but their abundance and diversity is still largely unknown. Not only are they very small, they are also very diverse. Some viruses look like space landers, encapsulated in strange protein costumes, others are spherical, thin or elongated. Some viruses have single stranded RNA as their hereditary material, while others carry their blueprint in double stranded DNA molecules. The one thing that is common to all viruses is that they have next to nothing in common. For instance, as they do not have any metabolism they lack unifying genes, like the ribosomal genes found in bacteria and eukaryotes. They are, essentially, a black box in the tree of life.
For most hosts, including my favorite ducks, we know next to nothing of the true virus diversity. We tend to study one host and one pathogen at the time, in isolation. But this is unwise, as most pathogens can infect more than one host species, and most hosts can harbor more than one pathogen species. Therefore we ignore putative important interactions between pathogens and hosts.
One research theme in the lab is to analyze pathogen assemblages in ducks. Last week we published a first paper on this in Infection, Genetics and Evolution; on the temporal dynamics, diversity and interplay of three viruses in Mallards. The viruses were: the influenza A virus, our standard virus in the lab; avian paramyxoviruses (AMPV-1), a less well-known genus, but which includes the poultry pathogen Newcastle disease virus; and avian coronaviruses (CoV). The latter is the least known of the bunch, but includes species that cause disease in domestic fowl, and are distantly related to viruses that cause respiratory infections in humans.
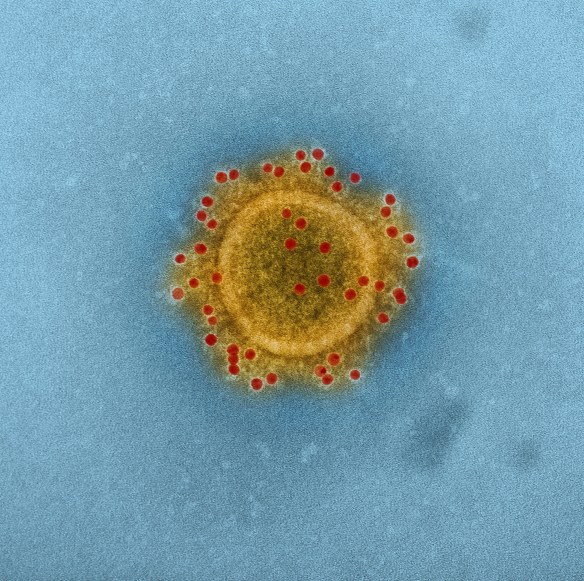
A Middle East Respiratory Syndrome Coronavirus – better known as MERS-CoV. This novel zoonotic pathogen is of great concern in the Middle East. Avian CoVs are not linked to disease in humans. (Photo NIAID)
From earlier studies, we know that these viruses circulate in the same population of ducks at the same time. Thus, one could imagine that they could compete over hosts, especially if they infect similar types of cells, or utilize similar receptors. Or, competition may be mediated by immune responses, where the responses raised towards one infection also impair infections with other viruses. Or the other way around: viruses could facilitate one another, where infection with one virus increases the fitness of the next virus, etc.
We followed a set of 144 Mallards across an autumn season, analyzing each sample collected at their first capture occasion, and at any other recapture occasions that followed. As this is a stopover site, recaptures of individuals are common, and the total number of samples analyzed was over three thousand. Thus for each day and individual, we ran three different real-time PCRs, one for each virus. When a sample was positive for a virus, we tried to cultivate it using egg inoculations, or sequenced part of its genome by targeted PCR assays.
There was a high prevalence of influenza A virus, comprising of 27 different subtype combinations, while APMV-1 had a comparatively low prevalence (with a peak of 2%) and limited strain variation. Avian CoVs were common, with prevalence up to 12%, and sequence analysis identified different putative genetic lineages. An investigation of the dynamics of co-infections revealed a synergistic effect between CoV and IAV, whereby CoV prevalence was higher given that the birds were co-infected with IAV. There were no interactive effects between IAV and APMV-1.
The diversity of viruses in these Mallard hosts is quite astounding, especially for CoVs and avian influenza viruses. With that many distinct variants circulating simultaneously in the population, the exposure much be very high to individuals. Entangling cause and effect in this system will ultimately depend on a combination of experimental and screening studies, but is a worthy goal. As for detection, recent advances in sequencing methods may open up broader studies on co-occurrences of viruses in hosts. We’ll see when, and how we can do that. Hopefully soonish.
Disease dynamics are the result of an interplay between parasites, host immune responses, and resources and it is imperative that we begin to include all factors to better understand infectious disease risk.
Wille, M., Avril, A., Tolf, C., Schager, A., Larsson, S., Borg, O., Olsen, B & Waldenström, J. 2015. Temporal dynamics, diversity, and interplay in three components of the virodiversity of a Mallard population: Influenza A virus, avian paramyxovirus and avian coronavirus. Infection, Genetics and Evolution 29: 129-137. [doi: 10.1016/j.meegid.2014.11.014]
*******************************************************************************************************************
If you enjoyed this post, or other posts on this blog, why not follow the blog via email, Feedly or get updates via Twitter by following @DrSnygg?


