By Jonas Waldenström
When I write these lines, the storm Simone is hitting southeast Sweden. The trees around our house seem to move erratically, trunks bend, and the few remaining leaves are shaking. Things twirl in the wind, and strange tweaking sounds are coming from the chimney. A night better spent indoors, safe and sound. Perhaps, like me, sipping on a nightcap, and pray you don’t wake up to a devastated neighborhood.
Apart from the storm – which the news says is THE WORST in ages (which they often say) – the rain and the compact darkness are hallmarks of the season. The transition from autumn to winter is concomitant with an increase in the prescription of antidepressants. People, each and own, sit in their cabins and vegetate and wait for the spring, and the return of life to our north latitudes. Most depressing to me is the farewell to birds. No more fluttering songbirds in the garden, no aerial insectivorous swifts, no foliage gleaners, just the odd wet, miserable Jackdaw hoping to get lucky with the garbage bin.

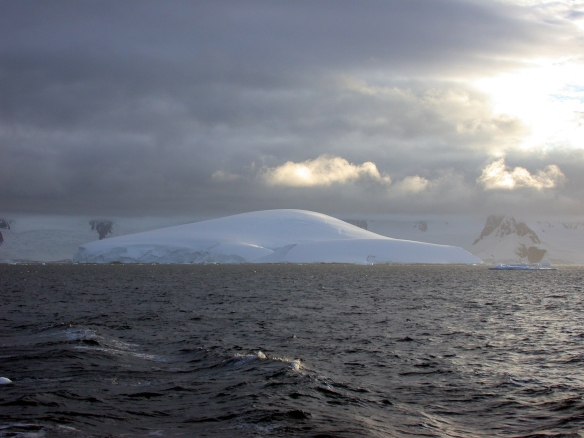
The aftermaths of Gudrun, the worst storm in recent times. It remains to see how the world looks tomorrow, after Simone.
It is also the Halloween season, or the All Saints’ Day as it is remembered for here in Europe. A time when we should be scared by gauls and ghosts, real or in costumes. But do you know what scares me the most? Truly. It is not the (imminent) zombie apocalypse, nor is it motor bikers with baseball bats – no it is tiny little organisms. You might have heard their names before: MRSA, ESBL, or NDM1.
There is a silent pandemic out there. Not the influenza crash-and-burn style pandemic. No, a silent one, a slow step-by-step walk towards the abyss. In a not too distant future (actually already today) some pathogenic bacteria will be resistant to the antibiotic drugs we use to treat infections. Widespread resistance changes medicine, it changes the way our healthcare function, and the options a patient has for treatments. It changes the outcome of infection, it causes deaths. For instance, multi-resistant tuberculosis strains are spreading that can cope with almost all the compounds normally used for treatment.
We don’t have too look far to find resistant bacteria – a trip to your local hospital is sufficient. In that ecosystem, we select for bacteria that can withstand drug therapy – the results are nosocomial infections, or hospital-acquired infections. It has been estimated that a fifth of all infections are nosocomial, meaning that this is an acute and severe problem for the health sector. Thinking of it, actually, you don’t have to go to the hospital to find resistant bacteria. Simply opening the door to the fridge or the freezer will get you there. An increasing proportion of the food items we eat, especially meat, but also vegetables, contain bacteria that carry resistance genes.
Bacteria have always evolved resistance to the chemicals used to fight them. We humans have waged war on bacteria for approximately a hundred years. Antibiotic compounds were an enormous success initially. Deadly infections were now made harmless and possible to control by prescription of drugs. When wound infections were treatable surgery became commonplace, and not a last resort. Not only appendicitis and cancers were treated, antibiotics opened the possibilities of today’s modern surgery where even organs can be moved between patients.
But what we didn’t think of in the early hay days of antibiotics was the vast evolutionary history of bacteria and chemical warfare. Although the human ingenuity is commendable, there is yet no compound invented that has not been tried in the evolutionary battle between microorganisms. Remember, fungi have tried to kill bacteria for billion of years. And through horizontal transmission between bacteria – sometimes across genera or families – genes evolved in one environment can find a home in a new bacterium and render the receiver new properties. A scientific niche the last few years have been to go searching for antibiotic resistance genes in preserved sediments, for instance in permafrost in Siberia, or in the deposits in caves or closed lakes. These studies have shown that, without a doubt, the genes have been around for a looooong time, thousands of years before humans even spoke the English language.
Actually a very poor representative of a scientific petri dish. In my lab that one likely was on its way to the autoclave. But you get it: bugs and gloves = dangerous
If you have read this far your either drunk or a masochist. My intention was to write about our latest article, exploring the occurrence of antibiotic resistance in wildlife in Chile, but the storm brought on the doom and gloom feelings. We’ll talk about solutions another night, when the storm winds are not howling across the land. But remember to be afraid – very afraid – of the slow pandemic that is about to change our way of life, as we know it.











